Species of the Pleurotus ostreatus complex (the oyster mushrooms) are one of the main groups of cultivated edible mushrooms in China. Due to phenotypic plasticity and morphological stasis, the species delimitation of the complex is often difficult and controversial. How to recognize the species of the complex rapidly and accurately is an urgent task in the mushroom industry.
Research group lead by Prof. YANG Zhuliang at Kunming Institute of Botany, Chinese Academy of Sciences (KIB/CAS) has recently conducted a study of the oyster mushrooms.
In this study, 284 strains from different mushroom spawn preservation centers, companies, and field isolations were investigated via phylogenetic analyses inferred from sequences of the internal transcribed spacer region (ITS), translation elongation factor 1-α (TEF1) and two genes that encode subunits of RNA polymerase ІІ (RPB1 and RPB2).
Data indicated that the 284 strains of the Chinese oyster mushrooms were divided into seven lineages, representing one morphological species each, namely P. abieticola, P. eryngii, P. cf. floridanus, P. ostreatus, P. placentodes, P. pulmonarius, and P. tuoliensis. The study revealed that 56 % of all the samples were labeled with improper scientific names. Meanwhile, RPB2 was indicated as the most promising candidate segment of DNA barcode for the P. ostreatus species complex.
This study has recently been published in Mycological Progress (https://link.springer.com/article/10.1007/s11557-016-1266-9).
It was supported by the National Basic Research Program of China (973 Program, No. 2014CB138305).
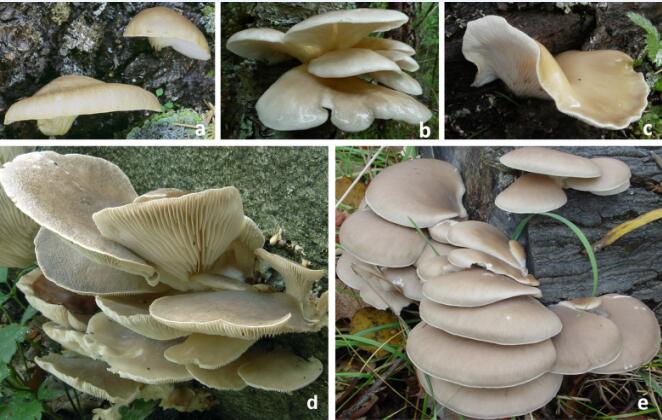

Fig. 1 Common species of the Pleurotus ostreatus complex in China: a. P. abieticola; b. P. pulmonarius; c. P. placentodes; d. P. ostreatus; e. P. cf. floridanus (Image by KIB)
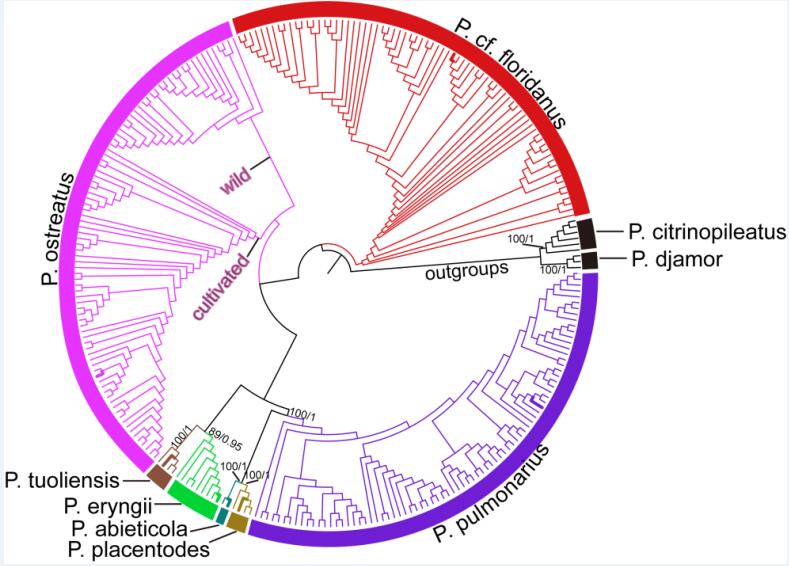
Fig. 2 Phylogenetic tree inferred from the nucleotide sequences of ITS-TEF1-RPB1- RPB2 (Image by KIB)
Contact:
Key Laboratory for Plant Diversity and Biogeography of East Asia, CAS
Kunming Institute of Botany, CAS
Prof. Dr. YANG Zhuliang
Email: fungi@mail.kib.ac.cn




