Glucose is the primary source of energy and substrate for cells, and its uptake through the cell plasma membrane is largely dependent on the glucose transport (GLUT) family.
GLUT1, one of the GLUT family members, is a ubiquitously expressed membrane protein. It is responsible for the constant uptake of glucose in many tissues.
Yan et al. reported the crystal structure of human GLUT1 at 3.2 angstrom resolution in 2014. However, the detailed distribution of GLUT1 on native cell membranes was not clear, and how protein organizations link to their physiological functions remained unknown.
With the modification and updating of cell membrane structure, many studies have found the heterogeneous and clustering distribution of membrane proteins.
A recent Protein Layer-Lipid-Protein Island (PLLPI) membrane model emphasized that a dense protein layer formed on the ectoplasmic side of the membrane and dispersed protein microdomains formed on the cytoplasmic side.
To address the distribution characteristics and the underlying organization mechanisms of GLUT1, super-resolution fluorescence microscopy has provided a particularly suitable tool. It breaks the diffraction barrier and achieves a lateral resolution in the tens of nanometers. It is very suitable for directly monitoring the size and stability of protein aggregations.
Prof. WANG Hongda of the Changchun Institute of Applied Chemistry, Chinese Academy of Sciences and Prof. XIONG Wenyong from the Kunming Institute of Botany, together with their team members, first investigated the distribution and assembly of GLUT1 at a nanometer resolution by direct stochastic optical reconstruction microscopy (dSTORM).
They found that the transporter formed clusters with an average diameter of ~250 nm on HeLa cell membranes. There was a precise spatial association between GLUT1 and lipid rafts, which resolved the debate surrounding the localization of the transporter in membrane domains.
Regarding the organizational mechanism of GLUT1 clusters, they revealed that not only can the lipid rafts’ environment stabilize their existence, but the actin cytoskeleton and N-glycosylation also play important roles in the clusters’ formation.
Moreover, they found that the activation of GLUT1 by sodium azide or MβCD did not increase its membrane expression but induced the decrease of the large clusters.
The results suggested a potential association between distribution and activation. Their teams’ work may form a step forward in our understanding of the molecular mechanism of GLUT clustering and glucose uptake.
This work has been published in a paper in PNAS entitled Mechanistic insights into GLUT1 activation and clustering revealed by super-resolution imaging. It was supported by the National Key R&D Program of China, the National Natural Science Foundation of China, and the Yunnan Provincial Science and Technology Department of China.
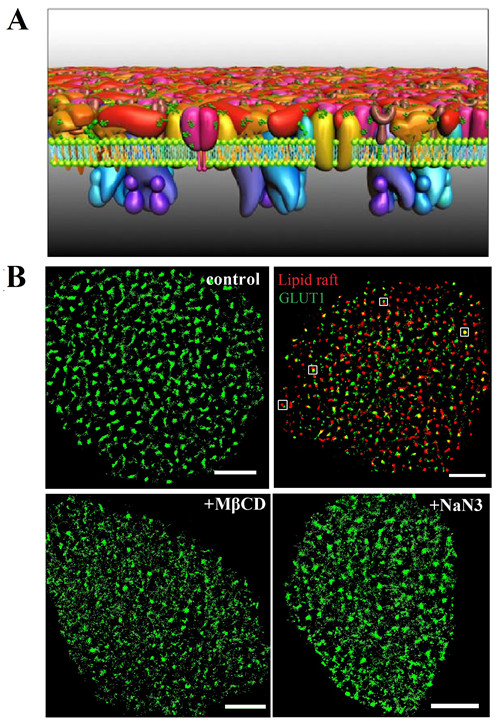
Fig. A) The PLLPI model of the cell membrane. B) dSTORM images of GLUT1 on the cell membrane; GLUT1 and lipid rafts; and GLUT1 distribution after MβCD and NaN3 treatment. Scale bars, 5 μm. (Image by WANG Hongda)
Contact:
Gao Jing
Changchun Institute of Applied Chemistry
E-mail: gaojing@ciac.ac.cn
(Editor:YANG Mei)




