Fragaria, commonly known as strawberry, is one of the most popular berry fruits and also one of the youngest domesticated plants.
The strawberry cultivated area of China ranks first in the world, and Yunnan is the main production area of perpetual-flowering strawberries. Southwest China is rich in wild strawberry germplasm resources, and has been recognized as an important distribution center of wild strawberry resources in the world.
The genus Fragaria has extensive interspecific hybridization and polyploidization, which makes the studies on the origin of organelle genomes and the evolution of nuclear-cytoplasmic interactions still largely unrevealed. This also hinders the effective evaluation and utilization of wild strawberry germplasm resources.
The formation and maintenance mechanism of genetic diversity is one of the essential questions in evolutionary genomics.
Compared with chloroplast genomes, plant mitochondrial genomes are generally characterized by complex and diverse structures, low substitution rate, frequent recombination, etc. It is considered as an important genetic system for studying the evolution of genome structure and function.
Previous studies have demonstrated that “plant mitochondrial DNA evolves rapidly in structure, but slowly in sequence”. However, it has also been suggested that there may be different selection pressure or mutation repair mechanisms between the coding and noncoding regions of the mitochondrial genome, resulting in the differences in the evolutionary rates.
To better understand the evolutionary features and mechanisms of plant mitochondrial genomes, researchers from the Kunming Institute of Botany of the Chinese Academy of Sciences (KIB/CAS) have assembled 13 complete mitochondrial genomes from Fragaria wild species and studied the sequence variation characteristics (substitution rate, indels, inversions) of their coding and non-coding regions.
The research team found the genome-wide nucleotide substitution rates are generally consistent between noncoding regions and synonymous sites in Fragaria mitogenomes, but indels and inversions occur more abundant in non-coding regions. Fragaria mitochondrial genomes experienced frequent rearrangements, so the genomic structure varies obviously even among closely related species.
This study indicates there are a large number of micro-inversion mediated multinucleotide substitutions (2-18 bp) in Fragaria mitochondrial genomes, and these continuous mutation sites adversely affect local mutation rates and phylogenetic signal. Moreover, a gain-and-loss model can explain the general lack of homology among plant mitogenomes.
This research was financially supported by the National Key Research and Development Program of China. Results have been published on New Phytologist entitled "Fragaria mitogenomesevolve rapidly in structure but slowly in sequence and incur frequent multinucleotide mutations mediated by micro-inversions".

Figure 1. Substitution rates in noncoding and coding sequences of Fragaria mitogenomes. (Image by KIB)
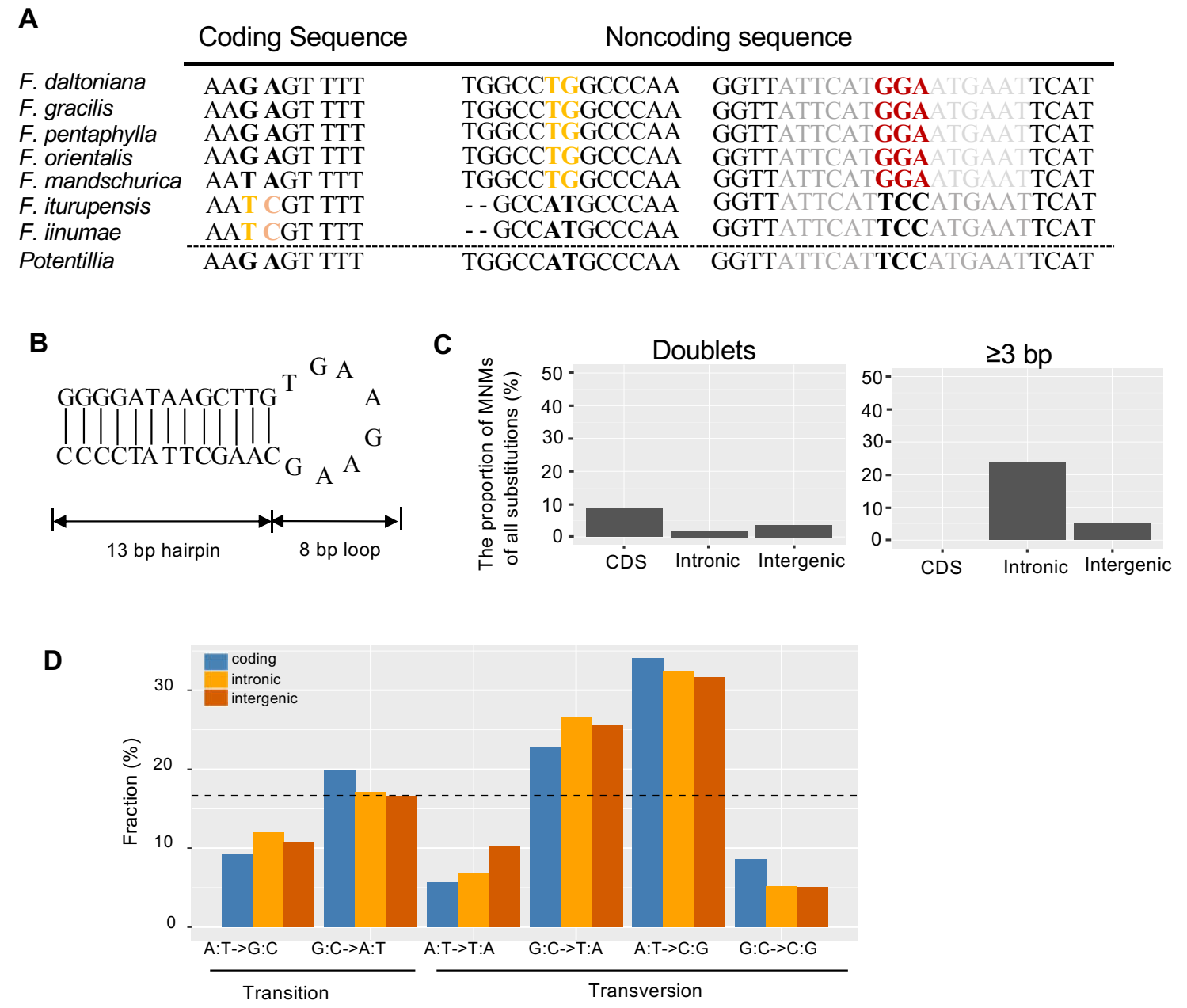
Figure 2. Multinucleotide substitutions and mutation spectra. (Image by KIB)
(Editor: YANG Mei)




